PDF Document Processing Pipeline with n8n
I got into the habit of browsing my LinkedIn feed on my smart phone. Every time I see a post about an interesting paper, I download it onto a cloud drive for later reading. Many of these papers were posted on arXiv.org, the open-access digital archive and distribution server for scholarly, non-peer-reviewed preprints (or postprints) in physics, mathematics, computer science, quantitative biology, quantitative finance, statistics, and related fields.
There are two challenges:
- the names of PDF files from arXiv.org mostly comprise numbers, making it difficult to find papers on the cloud drive;
- most papers do not include the actual reference of where they were published.
In this project I implemented a document processing pipeline with n8n (“n-eight-n”), a low-code workflow automation tool that enables users to connect applications, APIs, and databases to automate tasks. As an open-source, “fair-code” platform, it allows for self-hosting (via Docker), giving users full control over their data, privacy, and infrastructure.
This n8n workflow processes PDF files and changes the names of these files according to the title of the paper. In addition, it extracts the abstract and meta data of the papers and stores then for later processing. The workflow includes these five steps:
- load PDF files from staging folder and extract text and meta data,
- extract or generate the abstract of the paper,
- create a new filename from the title, add a hash string from the original filename to distinguish papers with the same title,
- move the original PDF file from the staging folder to the destination folder with the new filename, and
- save the metadata for analysis and further processing.
The diagram below shows the complete workflow. You can find the diagram and the complete code on the repo XXXX.
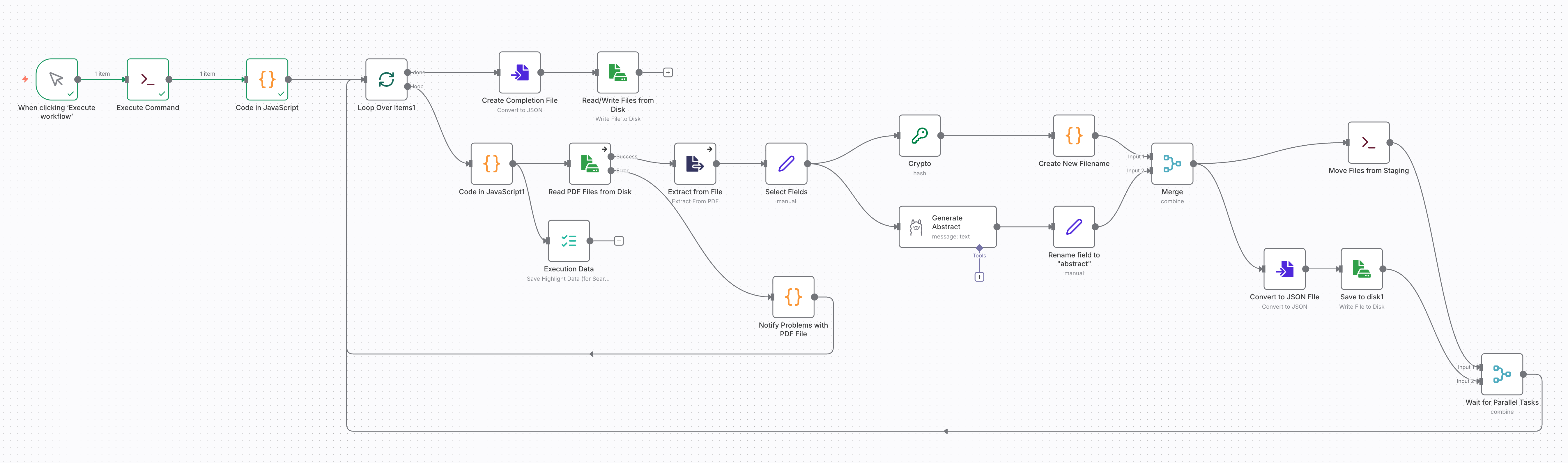
I implemented this project on a self-hosted, containerized n8n instance. You may also follow the steps to implement the workflow in the cloud version of n8n, though you need to change operations on the local file system with the respective nodes for cloud storage, such as Amazon S3.
First setup your n8n instance and make two folders available: one for staging and one for the final storage of the PDF documents and data files. In my project I used the directories /data/ai-papers-staging and /data/ai-papers within the container and mapped /data to a fold on the host file system.
Launch n8n and create a new workflow.
You start out with the trigger for the workflow. Choose “When clicking ‘Execute workflow’,” this will run the workflow to process all PDFs files in the staging folder. If you rather want your workflow to run continuously and process files as they get uploaded to the staging folder use the “Local File Trigger” with “Changes Involving a Specific Folder.”
You can load multiple files at once with the “Read/Write Files from Disk” node. However, this took a long time with just a bit over 100 files. Loading all at once before any further steps are processed also bears the risk of having to process everything all over again.
Instead, you build a workflow that pulls a listing of the files and then processes each PDF file in a loop. If a step fails and terminates the workfow, you will be able restart it and process the remaining documents.
There is no n8n node for listing files, so you have to create the file listing task in two nodes:
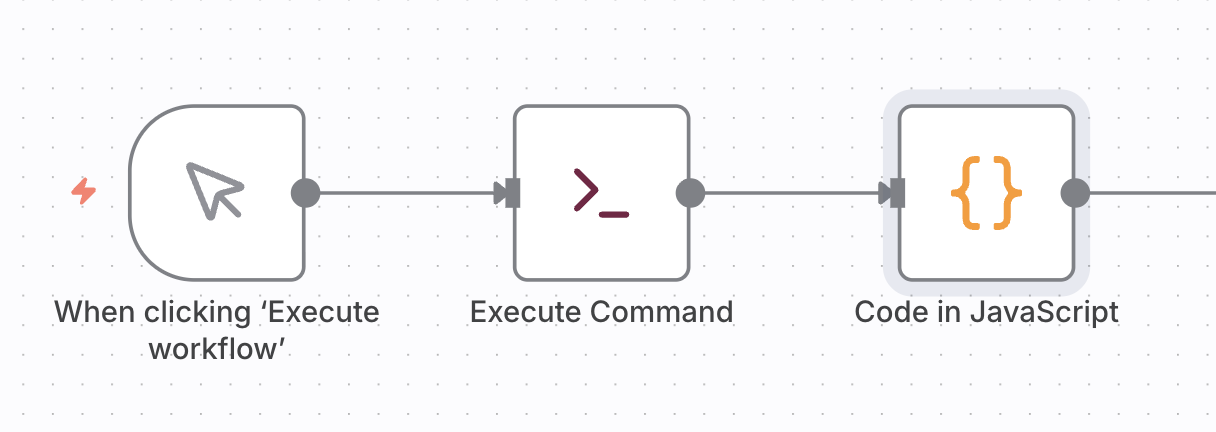
The first node, “Execute Command”, executes an shell command like
ls /data/ai-papers-staging/*.pdf
or
find /data/ai-papers-staging -name "*.pdf"
The second node, “Code in JavaScript” converts the file listing into JSON objects for the n8n data stream.
const stdout = $input.first().json.stdout;
const fileList = stdout.split('\n').filter(name => name.trim() !== '');
const numFiles = fileList.length;
// Map each filename to the standard n8n item structure
return fileList.map((fileName, index) => {
return {
json: {
fileName: fileName,
number: index + 1, // Current file position (1-based)
totalNumber: numFiles // Total count from the array length
}
};
});
The payload, that is carried throughout the workflow from one node to the other, is references as $item array. You only need to consider the first (and only) element, and extract the STDOUT content from the UNIX shell of the “Execute Command” node. The JavaScript code converts the multi-line text into a list of file names, and produces a new array of JSON objects to pass to the next node.
Enter the code snippets into the respective nodes and execute the workflow after you populated the staging folder /data/ai-papers-staging
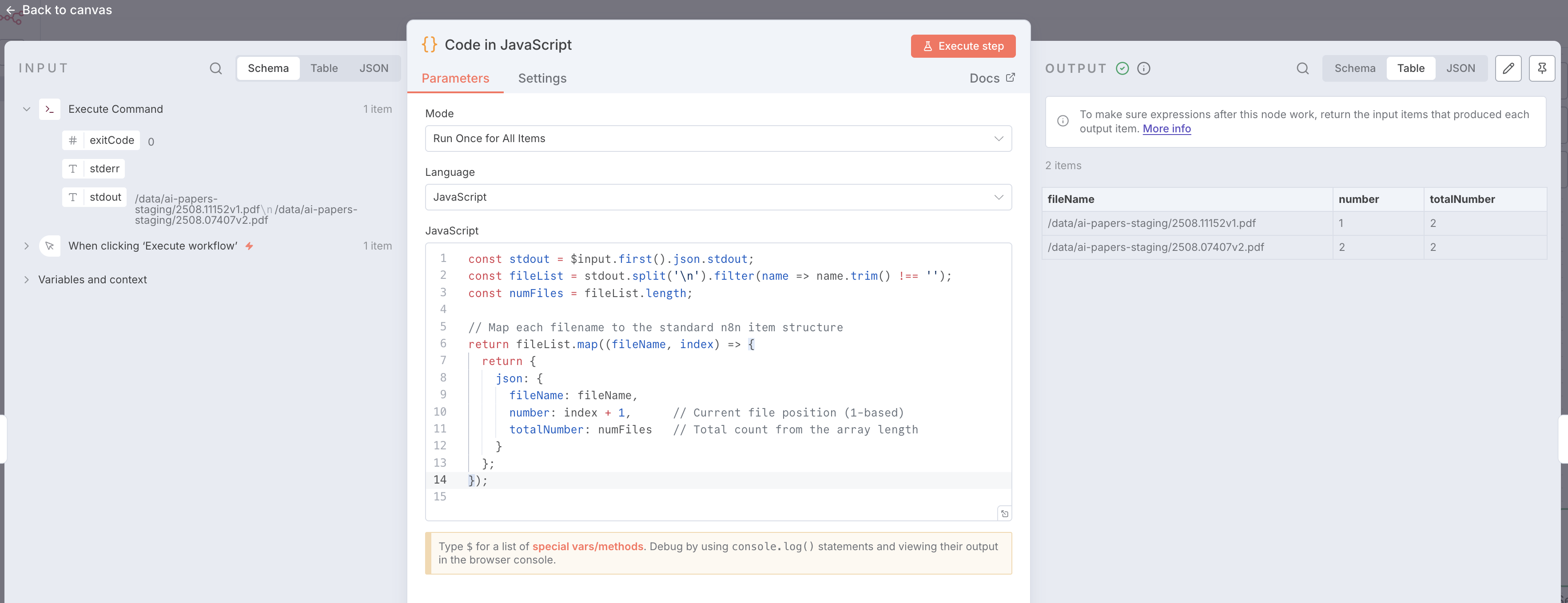
After the workflow has completed, double click on the “Code in JavaScript” node to see the results.
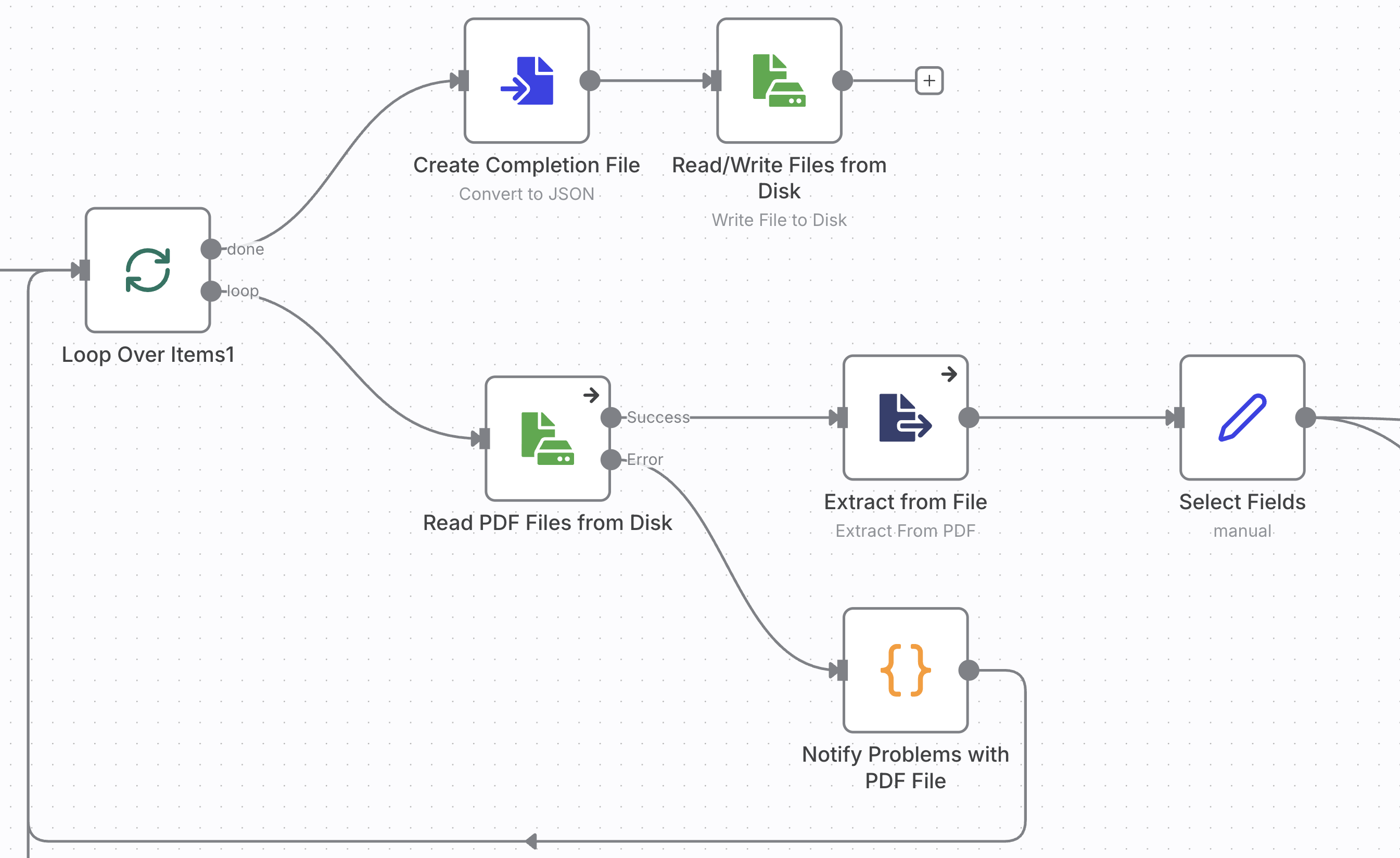
// Define the cleaning function first
function cleanString(text) {
if (!text) return "noname";
return text
.toLowerCase()
.replace(/[^a-z0-9]+/g, '_') // Replace special chars/spaces with _
.replace(/^_+|_+$/g, ''); // Trim underscores from ends
};
function generateHex(length = 8) {
let result = '';
const characters = '0123456789abcdef';
for (let i = 0; i < length; i++) {
result += characters.charAt(Math.floor(Math.random() * 16));
}
return result;
}
// Loop over input items
for (const item of $input.all()) {
// 1. Get the title from the JSON (falling back to 'untitled' if missing)
var title = item.json.Title || item.json.Title2 || "untitled";
// 2. Get the filename
var file_name = item.json.fileName || "";
file_name = file_name.replace(".pdf", "");
const hash = item.json.fileNameHash || generateHex(8);
// 3. Create the new filename
// We clean the combined string, then append .pdf
const baseName = cleanString(title.substring(0,24)) + "_" + hash;
item.json.NewFilename = baseName + ".pdf";
item.json.NewFileStem = baseName;
}
return $input.all();
In the body of the loop executes two independent tasks. The one on the top comprised of the “Crypto” and “Create New Filename” nodes forms a new filename for the document from its title and a hash of the original name. Most of the logic is in the code block. However, the build-in JavaScript does not provide a package for hashing alforithms. That’s why you need to use the “Crypto” node.
The branch below uses Ollama to generate an abstract from the text of the first page of the document.
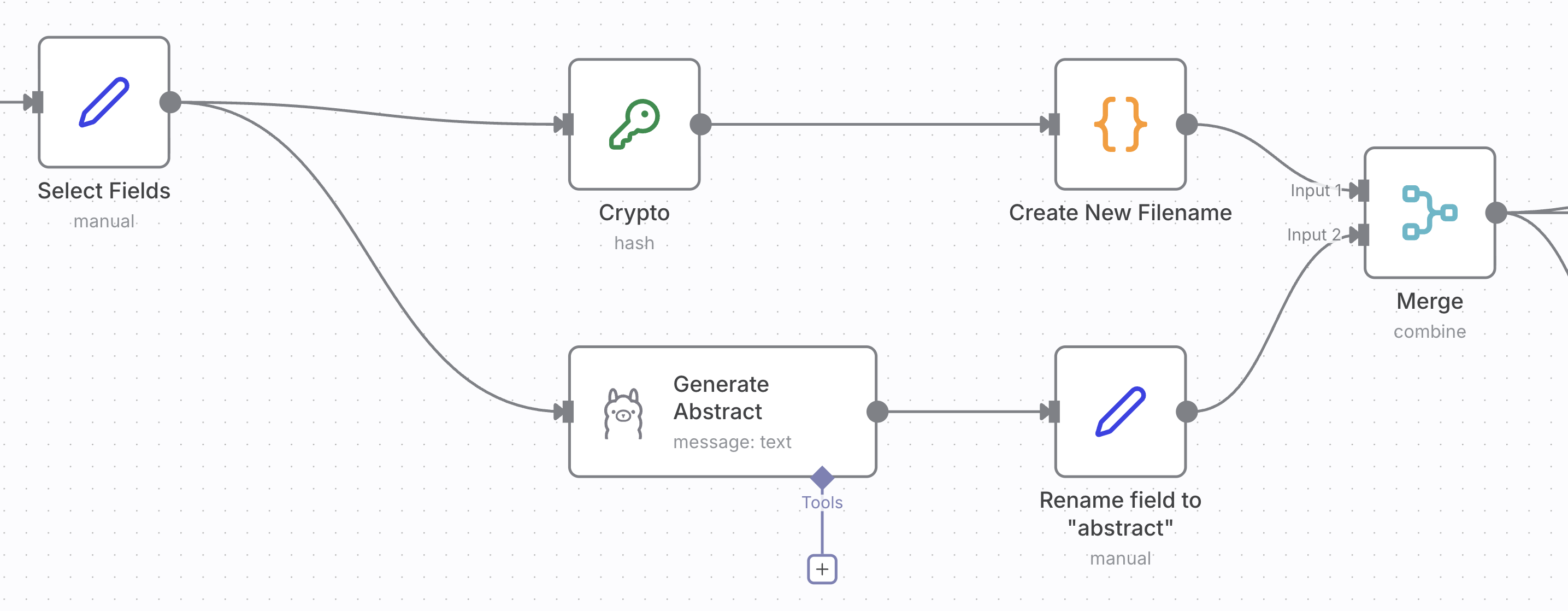
Both tasks create fields that you need to merged. Finally the data objects inserted into a data table, saved into a JSON file. The original file is renamed to the new name.
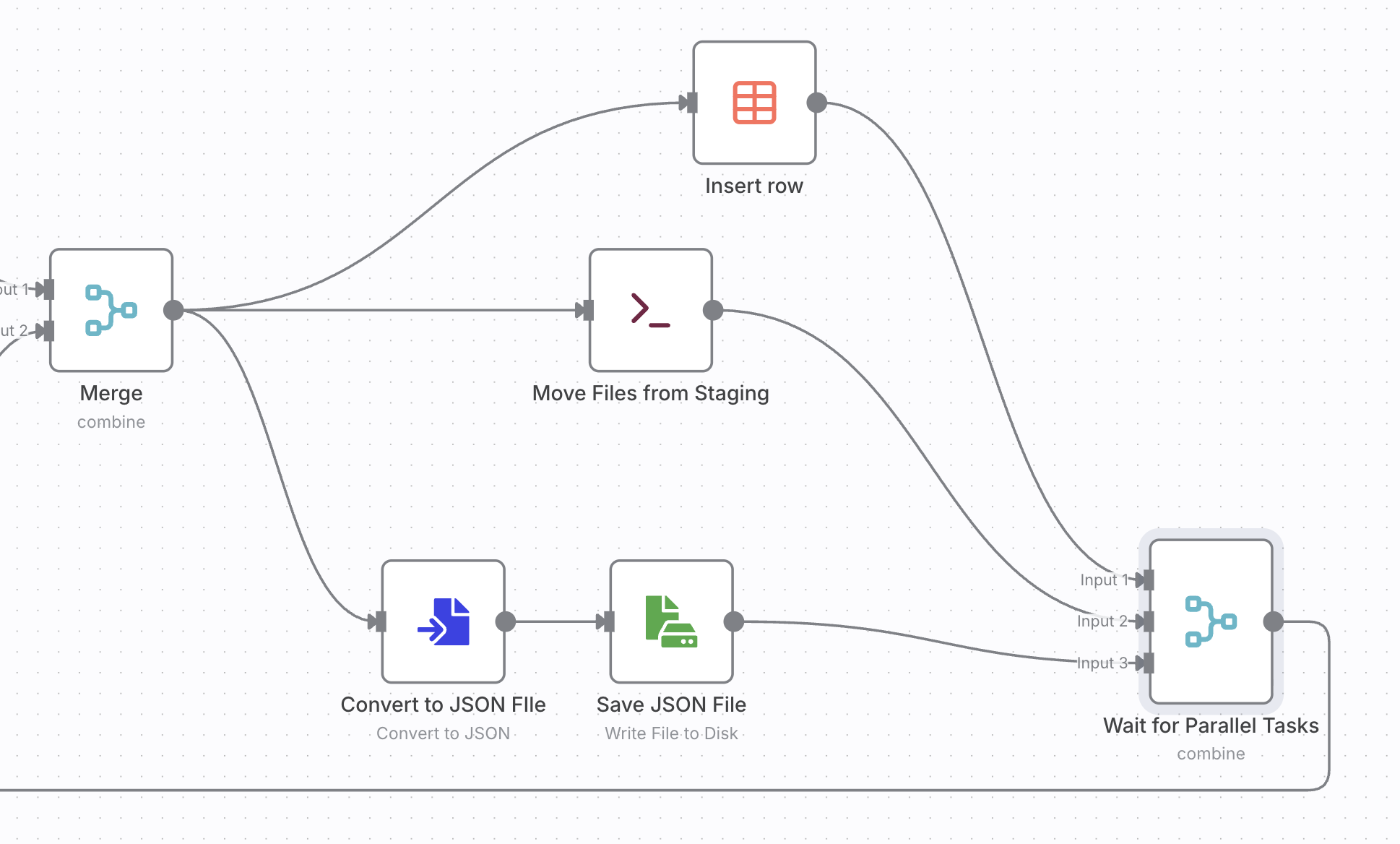
This workflow turned what used to be a messy, manual process into a structured and fully automated pipeline. By combining file system operations, metadata extraction, and lightweight AI processing in n8n, I now have a searchable, consistently named collection of research papers ready for further analysis. Beyond arXiv papers, the same pattern can automate academic literature curation, internal documentation management, or data ingestion for knowledge graph construction. It’s a simple example of how low-code automation can bridge the gap between research workflows and reproducible, scalable systems.